Research led by the University of Southampton has revealed the fundamental features of the SARS-CoV-2 coronavirus that causes COVID19. The researchers have produced the first model of a spike of the virus which shows how it disguises itself to enter human cells undetected, and the viral proteins which are the target of antibodies and vaccine research. The findings of this study could provide crucial information to help scientists currently searching for a vaccine.
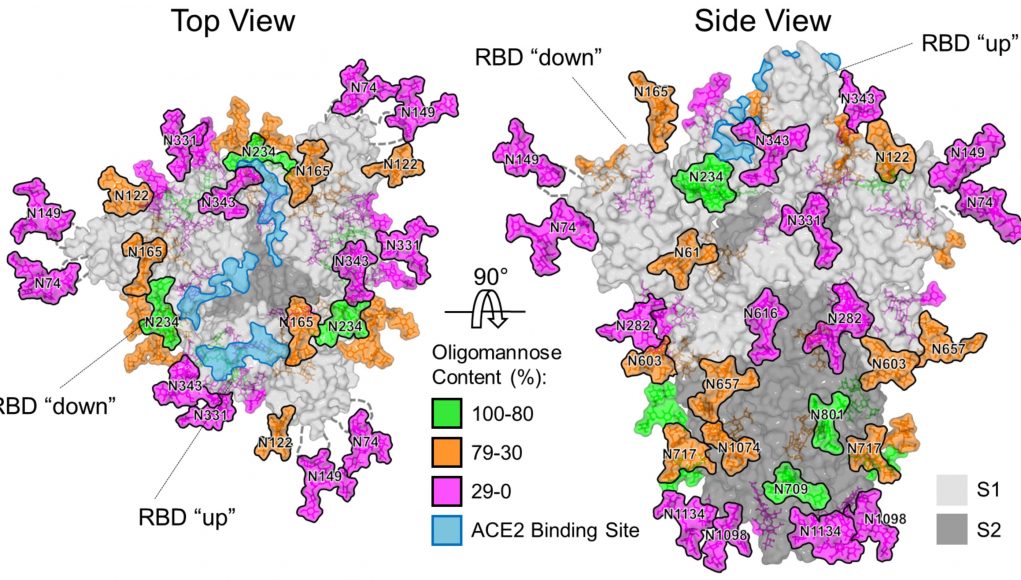
The SARS-CoV-2 virus has a large number of spikes sticking out of its surface which it uses to attach to and enter cells in the human body. These spikes are coated in sugars, known as glycans, which disguise their viral proteins and help them evade the body’s immune system.
The research team, led by Professor Max Crispin, studied the structure of the glycans covering the surface of a mimetic of a viral spike using equipment previously purchased through a grant from the Bill and Melinda Gates Foundation through the Collaboration for AIDS Vaccine Discovery.
They were then able to map the structure of the glycans which provides important information about how accessible the viral protein surface is to antibodies, this is an important step in vaccine design.
“By coating themselves in sugars, viruses are like a wolf in sheep’s clothing,” explained Professor Crispin. “But one of the key findings of our study is that despite how many sugars there are, this coronavirus is not as highly shielded as some other viruses.”
Find your dream job in the space industry. Check our Space Job Board »
“Viruses like HIV, which hang around in one host, have to evade the immune system constantly and they have a really dense coat of glycans as a shield to the immune system; but in the case of the coronavirus the lower shielding by sugars attached to it may reflect that it is a ‘hit and run’ virus, moving from one person to the next. However, the lower glycan density means there are fewer obstacles for the immune system to neutralize the virus with antibodies. So this is a very encouraging message for vaccine development.”
At Southampton, Professor Crispin’s team includes Ph.D. students Yasunori Watanabe and Joel Allen, and they worked closely with Jason McLellan’s team from the University of Texas who were the first to determine the structure of SARS-CoV-2. They have released their findings ahead of peer-review on the BioRxiv preprint server.
Professor Crispin’s team has a very strong history in analyzing the features of viruses and they have made key discoveries determining the features of the natively folded spike of HIV.
They are now working with partners who have developed candidate vaccines, including Professor Rogier Sanders at the University of Amsterdam, and are now analyzing the glycan content in Southampton. Evaluating the glycans on immunogens will determine how closely they mimic a natively folded viral spike and will help understand the immune response to vaccine candidates.
Provided by: University of Southampton
More information: Yasunori Watanabe et al. Site-specific analysis of the SARS-CoV-2 glycan shield. BioRxiv (2020). DOI: 10.1101/2020.03.26.010322
Image: Structure-based mapping of SARS-CoV-2 S N-linked glycans on its spike.
Credit: University of Southampton