Researchers have provided new insight on the mechanism underlying the control of gene expression in all living organisms, according to a study published today in eLife.
The findings, first reported on bioRxiv, could ultimately improve our understanding of how certain antibacterial drugs work against the enzyme RNA polymerase (RNAP) in treating conditions such as Clostridium difficile infections and tuberculosis.
Gene expression occurs when the information contained in DNA is used to produce functional gene products such as proteins and other molecules. The process has two stages. In the first stage, called transcription, RNAP reads the information in a strand on DNA, which is then copied into a new molecule of messenger ribonucleic acid (mRNA). In the second stage, the molecule then moves on to be processed or translated.
However, to help control gene expression levels, transcriptional pausing by RNAP can occur between the two stages, providing a kind of ‘roadblock’ where transcription may be terminated or modulated.
“A consensus pause sequence that acts on RNAPs in all organisms, from bacteria to mammals, halts the enzyme in an elemental paused state from which longer-lived pauses can arise,” explains senior author Robert Landick, Charles Yanofsky Professor of Biochemistry & Bacteriology at the University of Wisconsin-Madison, US. “As the fundamental mechanism of this elemental pause is not well defined, we decided to explore this using a variety of biochemical and biophysical approaches.”
Find your dream job in the space industry. Check our Space Job Board »
The team’s analyses first revealed that the elemental pause process involves several biological players, which together create a barrier to prevent escape from paused states. The process also causes a modest conformational shift that makes RNAP ‘stumble’ in feeding DNA into its reaction centre, temporarily stopping it from making RNA.
“We also found that transcriptional pausing makes RNAP loosen its grip and backtrack on the DNA while paused,” says Landick. “Together, these results provide a framework to understand how the process is controlled by certain conditions and regulators within cells.”
He adds that these insights could aid future efforts to design synthetic genes, for example to direct the pausing behaviour of RNAP in a way that yields desired outputs from genes. It could also help our understanding of how certain drugs, known as RNAP inhibitors, target the enzyme.
“For now, we would like to try and generate structures of paused transcription complexes obtained at a series of time intervals,” Landick concludes. “This would allow us to see exactly how parts of the enzyme move as it enters and leaves the paused state.”
Provided by:
eLife
More information:
Jason Saba et al. The elemental mechanism of transcriptional pausing. eLife (2019). DOI: 10.7554/eLife.40981
Image:
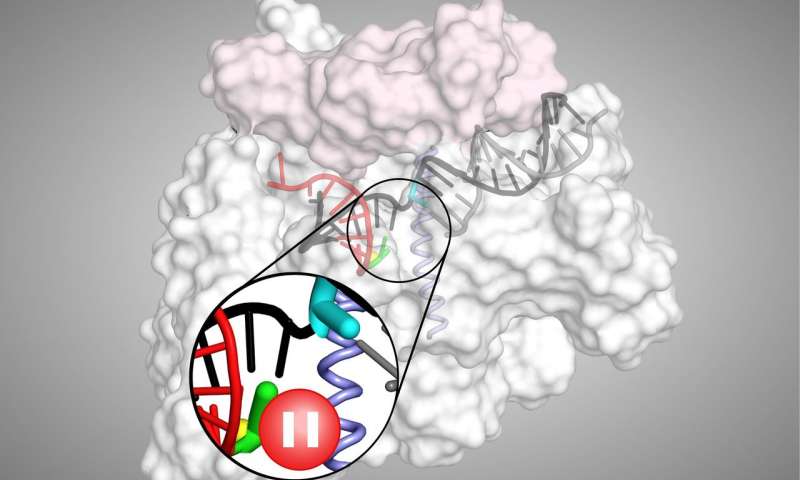
An illustration of the transcriptional pause process, which helps control gene expression in cells
Credit: Robert Landick