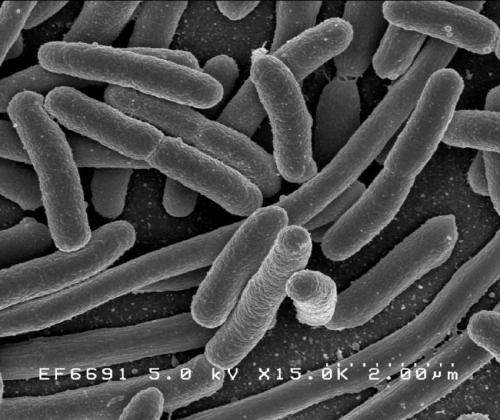
A genetic disruption strategy developed by University of Colorado Boulder researchers effectively stymies the evolution of antibiotic-resistant bacteria such as E. coli, giving scientists a crucial leg up in the ongoing battle against deadly superbugs.
These multidrug-resistant pathogens—which adapt to current antibiotics faster than new ones can be created—infect nearly 2 million people and cause at least 23,000 deaths annually in the U.S., according to data from the Centers for Disease Control.
In an effort to develop a sustainable long-term solution, CU Boulder researchers created the Controlled Hindrance of Adaptation of OrganismS (CHAOS) approach, which uses CRISPR DNA editing techniques to modify multiple gene expressions within the bacteria cells, stunting the pathogen’s central processes and thwarting its ability to evolve defenses.
“We now have a way to cut off the evolutionary pathways of some of the nastiest bugs and potentially prevent future bugs from emerging at all,” said Peter Otoupal, lead author of the study and a doctoral researcher in CU Boulder’s Department of Chemical and Biological Engineering (CHBE).
The CHAOS research is the culmination of work that began in 2013, when Otoupal and his colleagues began searching for genes that could act as a cellular kill switch for E. coli. When the scientists tweaked one gene at a time, the bacteria could adapt and survive. But when they altered two or more genes at once, the cell got weaker.
Find your dream job in the space industry. Check our Space Job Board »
“We saw that when we tweaked multiple gene expressions at the same time—even genes that would seemingly help the bacteria survive—the bacteria’s fitness dropped dramatically,” Otoupal said.
The CHAOS method takes advantage of this effect, pulling multiple genetic levers in order to build up stress on the bacterial cell and eventually trigger a cascading failure, leaving the bug more vulnerable to current treatments. The technique does not alter the bug’s DNA itself, only the expression of individual genes, similar to the way a coded message is rendered useless without the proper decryption.
“You can think of it in terms of a series of escalating annoyances to the cell that eventually cause it to weaken,” said Anushree Chatterjee, senior author of the study and an assistant professor in CHBE. “This method offers tremendous potential to create more effective combinatorial approaches.”
Although E. coli has nearly 4,000 individual genes, the exact gene modification sequence appears to matter less than the sheer number of genes that are disrupted, Otoupal said. Still, the researchers plan to continue optimizing the CHAOS method to seek out the most efficient disruptions.
The findings are outlined today in the journal Nature Communications Biology and could open new research avenues on how to best restrict a pathogen’s antibiotic resistance.
“Diseases are very dynamic, so we need to design smarter therapies that can gain control over their rapid adaptation rates,” Chatterjee said. “The emphasis in our lab is demonstrating the efficacy of these methods and then finding ways to translate the technology to modern clinical settings.”
“In the past, nobody really considered that it might be possible to slow down evolution,” Otoupal said. “But like anything else, evolution has rules and we’re starting to learn how to use them to our advantage.”
Provided by:
University of Colorado at Boulder
More information:
Peter B. Otoupal et al. Multiplexed deactivated CRISPR-Cas9 gene expression perturbations deter bacterial adaptation by inducing negative epistasis. Communications Biology (2018). DOI: 10.1038/s42003-018-0135-2
Image:
Escherichia coli
Credit: Rocky Mountain Laboratories, NIAID, NIH